
MalVac Tutorial Page

 |
|
 |
|
MalVac is an information resource for the Malarial Vaccine Candidates. It includes curated information for currently known Malarial Vaccine Candidates obtained from careful literature search. Apart from this MalVac also contains a detailed analysis of predicted Vaccine Candidates from three plasmodium species- P.falciparum, P.vivax and P.yoelii that have a probability of being involved in the adhesion of Plasmodium pathogens to the host cells. This probability is decided on the basis of MAAP Score. The analysis includes various computational approaches to identify the number of paralogs, orthologs, number of transmembrane helices, signal peptides, beta wraps and topology of the protein . It also provides information about conserved domains present in the Vaccine Candidate and blastp results reflecting similarity to host proteins. MalVac also contains detailed epitope information for all the selected Vaccine Candidates. It contains the linear B cell epitope data obtained from ABCpred and Bcepred servers and conformational B cell epitope data from CEP and Discotope servers. |
Salient Features of MalVac
|
| Query Making Options The informations present in MalVac Database can be retrieved through the "Database Search" tab which takes user to the MalVac Query Page. The information to be retrieved through this page can be divided into two layers. First Layer having general information of motif, topology, location, homology and antigenic regions and Second Layer having epitope and allergen informations for the vaccine candidates. The information present in MalVac Database for the First Layer can also be retrieved through the "Search Tools" tab which takes the users to the MalVac Advanced Search page. Through this page users can filter their results on the basis of Protein length, number of transmembrane spanning regions, localization and reliability class, presence or absence of betawraps, paralogs, orthologs, hits to Conserved Domain Database and Human Reference proteins. (I) Query Making Through "MalVac Query Page"(i) Select a species from the pull down menu list.  |
(ii) Provide an Orf ID corresponding to the selected species. For example if user selects Plasmodium falciparum, then the Orf ID provided by the user must correspond to Plasmodium falciparum. Multiple comma separated ORF IDs can also be provided eg- PFL0935c,PFL1960w,PF08_0142. This field is optional for first layer data retrieval but must be provided to fetch second layer data. Second layer data can be retrieved corresponding to a single Orf ID against the species selected.  |
(iii) The user can also retrieve the First Layer information corresponding to a keyword. This field is optional.
 |
(iv) Select checkbox corresponding to the data user wants to retrieve in Data Available frame.
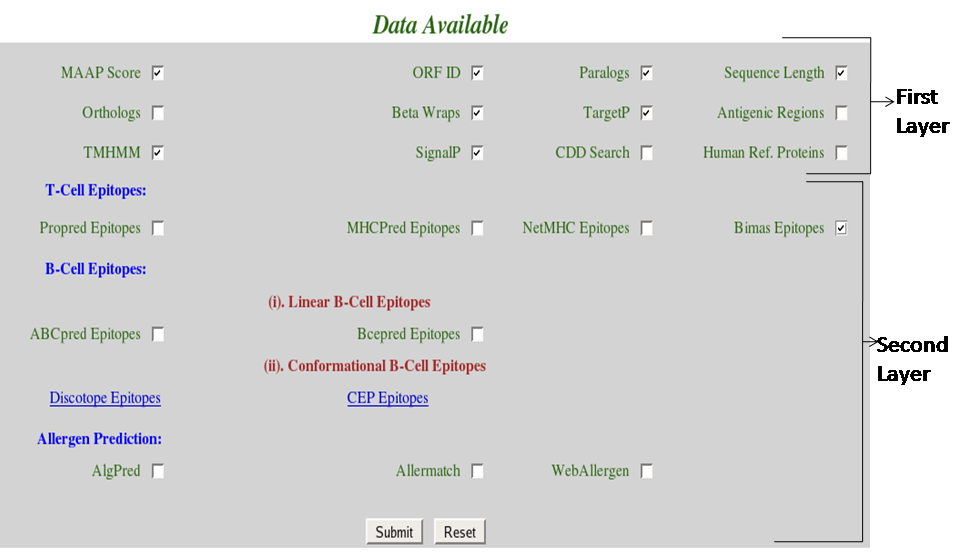 |
| (v) In the Second Layer the Conformational B-Cell Epitope data is available as flat files, these are hyper linked. The flat files can be saved by the file "save as" option by the users for future use.
(vi) Finally click the submit button. This takes users to the result page.  |
(vii) In the result page the entire quiried data can be saved by the user on clicking the hyperlink in blue "Export data for First Layer" which provides facility to save the First Layer Data in text file and the successive epitope and allergen hyperlinks "Export Bimas Epitope Data" etc which provides facility to save the Second Layer Data from individual servers in text files. 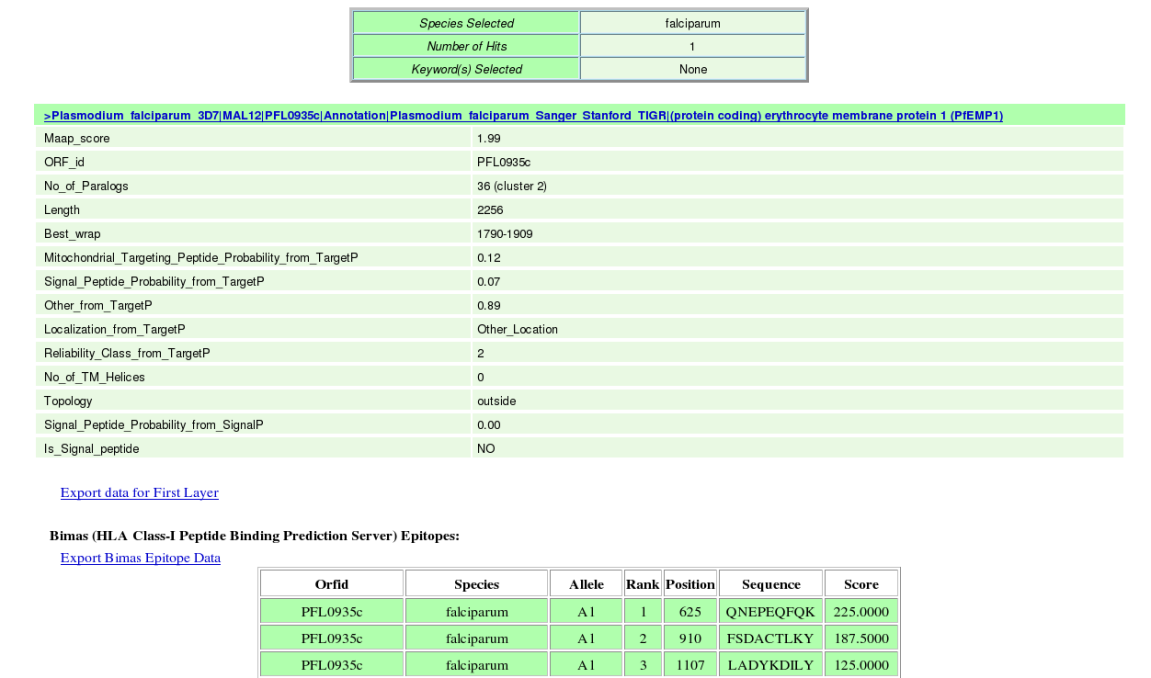 |
| (II) Query Making through "MalVac Advanced Search" page |
(i) Select a species from the pull down menu list. |
(ii) Select checkbox corresponding to the data user wants to retrieve in "Show information for:" frame |
(iii) Set the filters corresponding to the data selected through the drop down boxes.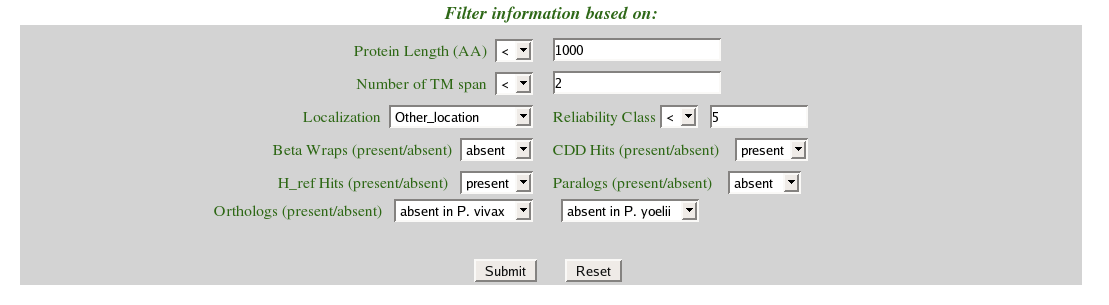 |
(iv) Finally click the submit button which takes users to the result page.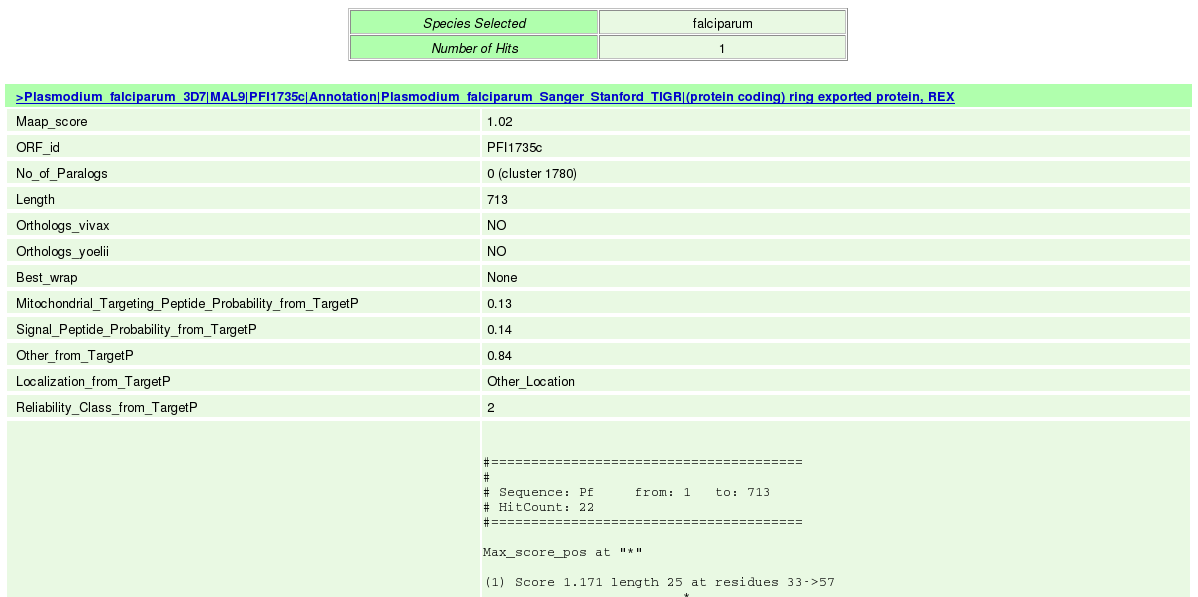 |
| (v) In the result page the entire queried data can be saved by the user on clicking the hyperlink in blue "Export" which provides facility to save the entire queried First Layer Data into a text file. |
| Viewing Known Vaccines Candidate Information
(i) The Malarial Known Vaccine Candidate Data can be viewed by clicking the "Known Vaccines" tab. The entire available list can be saved by the user into an excel sheet by clicking the hyperlink "Click to Export". |